|
A colormap is just a list of colors – 64 will normally do – which change smoothly from from one colour to another. is white, is black, and is a frightfully iridescent red. MATLAB stores colours as 1×3 vectors, where each element in the vector is the proportion of red, green, or blue, respectively. MATLAB’s colormap mechanism is just simple enough to be confusing. (More precisely, it is not a property of the MATLAB axes.) When you change the colormap, you change the colors of every datapoint in the image. Here’s the problem: the colormap is a property of the figure, not the data. No luck! What’s more, setting the colormap between calls to image() and imagesc() also doesn’t work. Hh = image(varargin,'CDataMapping','scaled') There is no ColorMap property on the Image class. We want to be able to highlight the regions in some colour other than grey. If we set 0.5*mask as the AlphaData property, then the colour we add will be at half transparency and the white areas will be fully transparent.īut this isn’t a very pleasant image. 0.5*mask is a matrix which is 0.5 everywhere that mask is 1 and 0 everywhere else. We want the image to be fully transparent where mask is 0 – so as not to fog out the underlying image – and partially transparent where mask is 1. If we did imagesc(mask), we would end up with grey everywhere and white only where we hoped to highlight – the opposite effect from the one that we sought.ĪlphaData is a property which sets the transparency of the image. 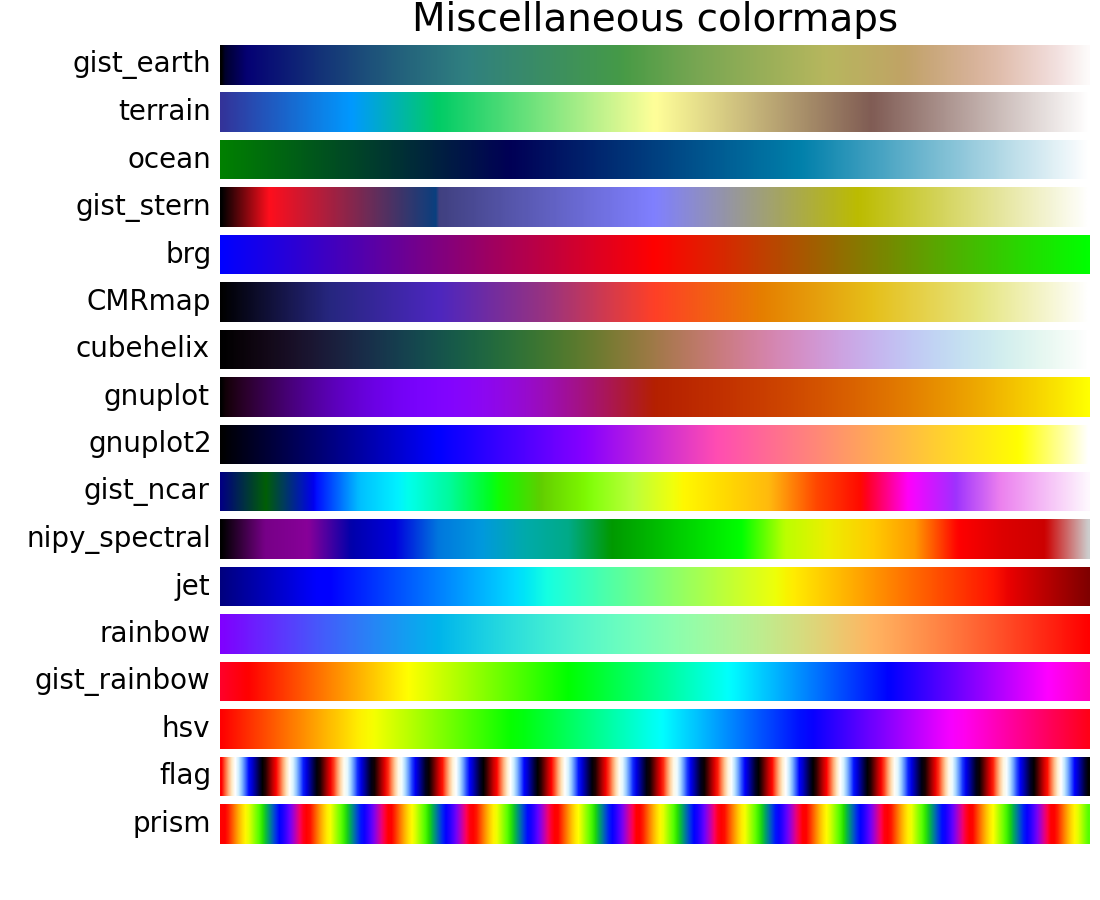
But we use imagesc(1-mask) because the gray colormap displays black at 0 and white at 1. Mask is a matrix which has a one at all the pixels that we want to highlight. Now, let’s load up a set of residues and try to overlay them on top of the first image: > resn=dlmread('1hel.resn') Let’s have a look at an example: > dists=dlmread('1hel.distances') The trouble is that I like to highlight parts of them, and that’s not straightforward in MATLAB. I need to make a lot of 2-D plots of protein distance matrices. However, there was issue of continued perplexity, and I’m not referring to why MATLAB insists on shouting itself at you. I realized that life’s too short to spend it tickling gnomes – especially when only one of them knows how to do linear regression, but he won’t tell you your p value unless you give him the right kinds of treats. I fired up MATLAB and I haven’t looked back. 
Soon, I found myself engaged in a reassessment of my life choices. But all that aside, I resolved, joining the group, to put aside my misgivings and give the gnomes another try. My overriding emotion, working in R, has been incomprehension: incomprehension at the gallery of ugly gnomes that populate the namespace and worried puzzlement over the strange incantations required to get them to dance in a statistically harmonious way.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
